Faculty Profile: YINGZE ZHANG, PhD
Dr. Yingze Zhang’s Research
Dr. Zhang’s current research is focused on the molecular and genetic basis of advanced lung diseases including COPD and ILD and biomarker discovery of these diseases. Dr. Zhang and her colleagues performed GWAS on idiopathic pulmonary fibrosis (IPF) and discovered several genes for predisposition, disease progression, and mortality in IPF. She is the co-lead author of their IPF GWAS study and a study demonstrated MUC5B gene as a disease outcome genetic marker published in the LANCET Respiratory Medicine and JAMA, respectively. Dr. Zhang is also the lead author for the validation of MUC5B as a predisposition gene for IPF reported in NEJM. Currently her lab is following up on the functional significance of IPF associated genes and genetic polymorphisms. Dr. Zhang and her colleagues also identified multiple blood protein biomarkers for disease progression and mortality in IPF. Her laboratory has identified the genetic basis for one of the elevated biomarkers, MMP7.
Dr. Zhang is one of the key investigators for the Pittsburgh COPD SCCOR study and directs the biorepository, biomarker discovery and genetic studies. She and her colleagues have reported whole genome sequencing for 132 subjects in the SCCOR cohort and identified rare genetic variants either predispose to or protect from the development of COPD. Her lab is following up on the functional significance of the COPD associated genes and genetic polymorphisms. She is a co-PI for a NHLBI funded study to identify blood biomarkers and to develop models based on the identified blood biomarkers in combination with clinical, demographic, radiographic and physiologic parameters for the prediction of long-term lung function decline in healthy smokers.
In addition to her studies in IPF and COPD, Dr. Zhang’s group also conducted functional analysis of genetic variants associated with bone mineral density. Her lab focused on one of the genes involved in the Wnt pathway, FZD1, and identified multiple transcription factors regulating FZD1 gene in an allele specific manner. The ultimate goal is to study these genes in a cohort of COPD since osteoporosis is more common in patients with COPD and further analysis the functional consequence of associated variants in lung and bone tissues of these patients.
Dr. Zhang also directs the Translational Research Core Lab (TRCL) for the Division of Pulmonary, Allergy and Critical Care Medicine and the Biobank for the Division of Cardiology and Vascular Medicine Institute (VMI). The TRCL has banked clinical samples from patients with different lung and vascular diseases since 2002 and has been served as a vital resource for translational research in PACCM, Cardiology and VMI.
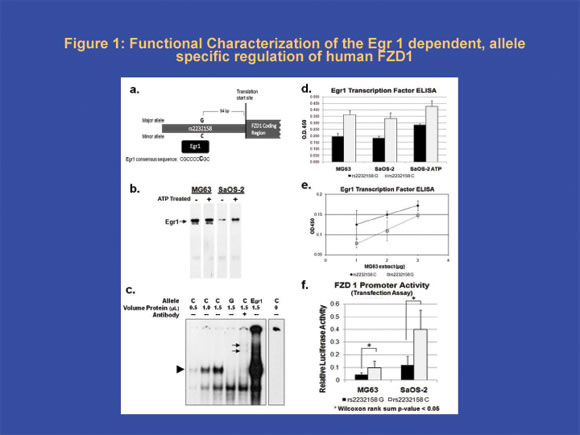
Functional analysis of the Egr-1 dependent/allele specific regulation of the FZD1
This figure illustrates the identification of Egr1 dependent, allele specific regulation of the FZD1 gene associated with rs2232158 polymorphism. a). Schematic of the FZD1 5’ UTR region and the creation of an Egr1 binding site associated with the rs2232158C allele. b) Western blot analysis of Egr1 protein in MG63 and SaOS-2 cells, the cell types for the in vitro study. c) Allelic specific binding to the rs2232158 specific probes with recombinant Egr 1 using EMSA. Increasing Egr1 specific complex (see arrowhead) was detected for the C allele with increasing Egr1 recombinant protein concentration. The Egr1 specific binding was abolished with Egr1 antibody treatment and super shift bands were observed (see arrows). d) Binding affinity for Egr1 to rs2232158 alleles. Transcription Factor ELISA (Panomics) and biotinylated probes designed for each allele of rs2232158 (G allele in black and C allele in white) were used. Differences in binding were tested using the Wilcoxon rank sum test (<0.05 indicated by *). e) Binding affinity for Egr1 to rs2232158 alleles at various concentrations of nuclear extract (G allele in black and C allele in grey). f) Activity of the rs2232158 specific FZD1 promoter. The promoter activity for different rs2232158 alleles were assessed by transfecting MG63 and SaOS-2 cell lines with luciferase reporter plasmids (G allele in black and C allele in white). Differences in activity were tested using the Wilcoxon rank sum test (<0.05 indicated by *)
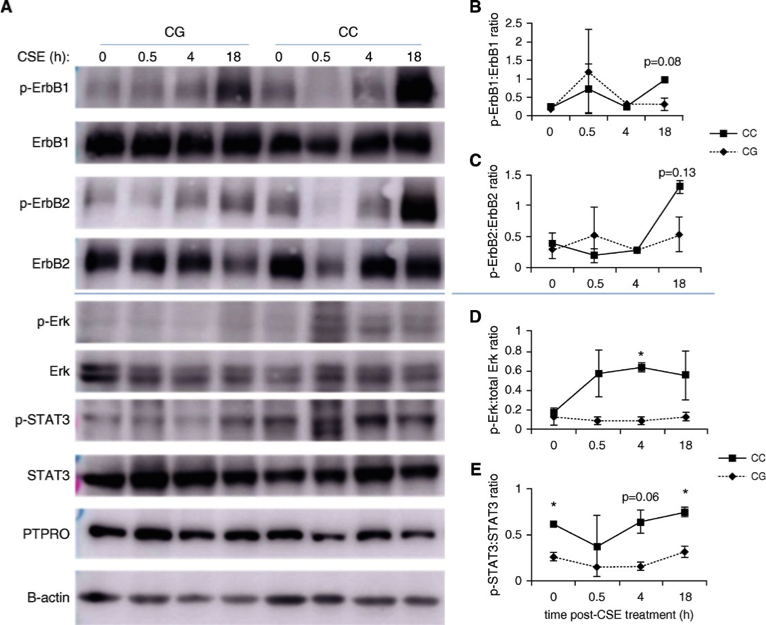
Figure 2. PTPRO rs61754411 genotype-specific intracellular signaling necessary for cell survival and proliferation after CSE and EGF treatments of human primary human bronchial epithelial cells. Human bronchial epithelial cells from individuals with no lung diseases bearing the PTPRO rs61754411-CG or -CC genotype (n = 2 of each) were cultured in air–liquid interface and treated with CSE for the times indicated (A–E). (A) Whole-cell lysates from treated cells were blotted for the EGFR subunits ErbB1 and ErbB2 and for the downstream signaling molecules Erk and STAT3. Blots above the blue line are representative of one unique individual, and blots below are representative of the other unique individual. As controls, PTPRO and β-actin were also analyzed. (B–E) Densitometry analysis (ratio of p-Akt to total Akt) was performed on these results. Data are presented as mean ± SEM. These same cell lines were cultured in monolayer and treated with recombinant human EGF for the times indicated (F–I). (F) Whole-cell lysates from treated cells were blotted for the EGFR subunit ErbB1 and for the downstream signaling molecules Erk and STAT3. Similarly, PTPRO and GAPDH were also analyzed as controls. (G–I) Densitometry analysis was performed on the blots shown in F for two unique individuals with PTPRO rs61754411-CC and -CG genotypes. Values are presented as a relative ratio of p-Akt to total Akt band intensity compared with those observed in basal, untreated conditions. *P < 0.05. For both treatments, the cells were starved with growth media without growth factor supplements for 4 hours before the treatment in growth media with reduced growth factor supplements (10% of normal culture conditions). Akt = protein kinase B; CSE = cigarette smoke extract; EGF = epidermal growth factor; EGFR = EGF receptor; ErbB1/2 = EGFR/human epidermal growth factor receptor (HER2); Erk = extracellular signal regulated kinase; GAPDH = glyceraldehyde 3-phosphate dehydrogenase; p-Akt = phosphorylated Akt; p-Erk = phosphorylated Erk; p-STAT3 = phosphorylated STAT3; PTPRO = protein tyrosine phosphatase, type O; STAT3 = signal transducer and activator of transcription 3.
Am J Respir Crit Care Med, 2017. Josiah Radder; Yingze Zhang; Alyssa Gregory; Shibing Yu; Neil J. Kelly; Joseph K. Leader; Naftali Kaminski; Frank C. Sciurba; Steven D. Shapiro; Am J Respir Crit Care Med 196, 159-171. Copyright © 2017 by the American Thoracic Society.
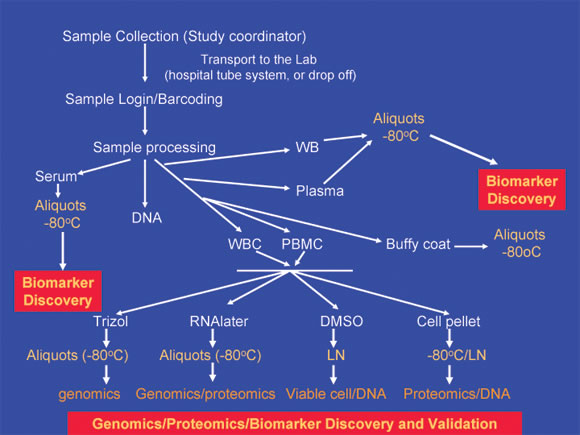
Schematic description of The translational Lab sample collection, processing and archiving.
This flow diagram outlines the sample collection, processing and archiving of clinical samples for the Translational Core Lab. The archived samples provide a vital resource for proteomics, genomics, and biomarker discovery and validation research.
Division of PACCSM
Academic Offices
NW 628 UPMC Montefiore
3459 Fifth Avenue
Pittsburgh, PA 15213
412-692-2210
Patient Care and Referral
Comprehensive Lung Center (CLC)
Falk Medical Building, 4th Floor
3601 Fifth Avenue
Pittsburgh, PA 15213
412-648-6161